Long non-coding RNAs (lncRNAs) have important biological functions and can be used as prognostic biomarkers in cancer. To identify a lncRNA prognostic signature for head and neck squamous cell carcinoma (HNSCC).
Researchers at the Shanghai Jiao Tong University School of Medicine analysed RNA-seq data derived from the TANRIC database to identify a lncRNA prognostic signature model using the orthogonal partial least squares discrimination analysis (OPLS-DA) and 1.5-fold expression change criterion methods. The prognosis prediction model based on the lncRNA signatures and clinical parameters were evaluated using the 5-fold cross validation method.
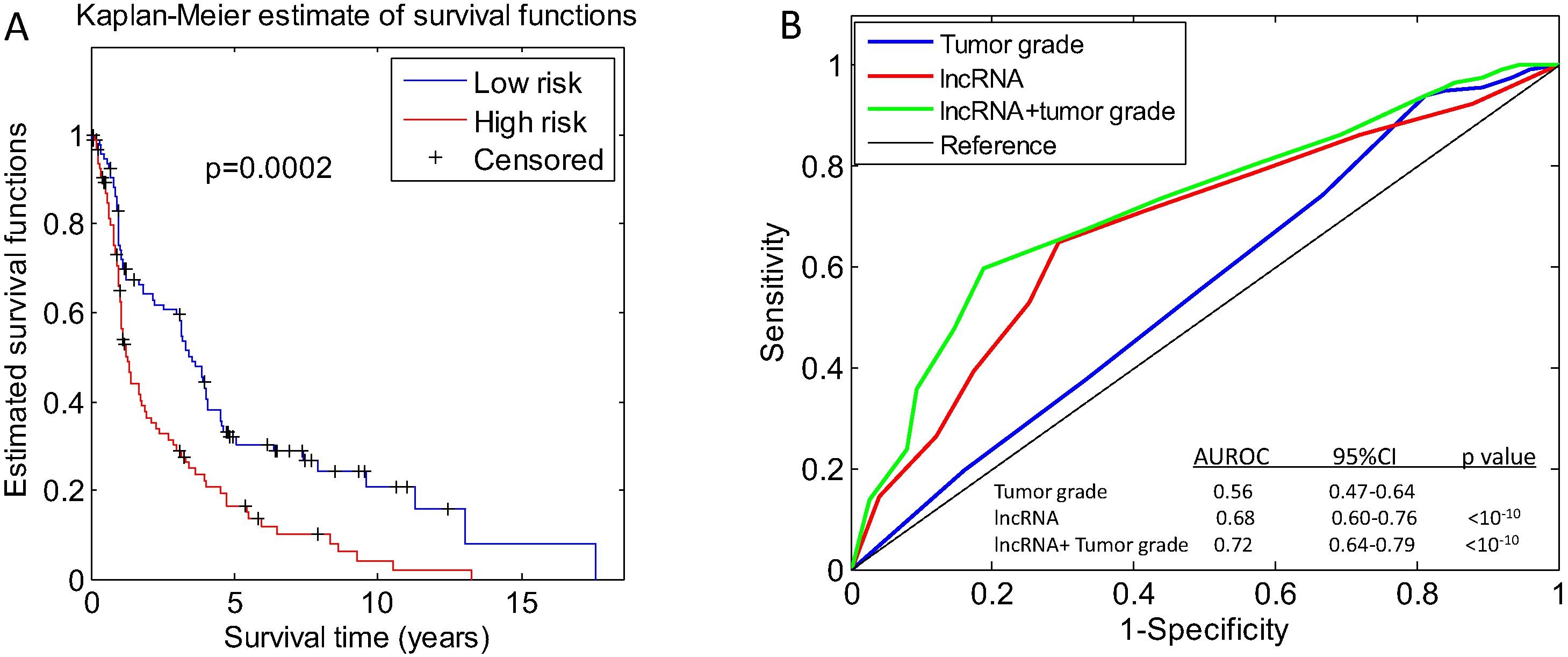
A total of 84 out of 3199 lncRNAs were significantly associated with the survival of patients with HNSCC. Using the OPLS-DA and 1.5-fold change selection criterion, 5 lncRNAs (KTN1-AS1, LINC00460, GUSBP11, LINC00923 and RP5-894A10.6) were further selected. The prediction power of each combination of the 5 lncRNAs was evaluated through the receiver operating characteristic (ROC) curve and a three-lncRNA panel (KTN1-AS1, LINC00460 and RP5-894A10.6) achieved the highest prognostic prediction power in the cohort. The patients were categorized into high- and low-risk groups based on their three-lncRNA profiles. Patients with high-risk scores had worse overall survival than those with low risk scores in the cohort. Multivariable Cox regression analyses showed that the lncRNA signature and tumour grade were independent prognostic factors for patients with HNSCC.
(A) The categorization of the patients based on the profiles of the three-lncRNA panel is associated with their survival time. (B) The prediction capability of the models constructed on the three-lncRNA panel and tumour grade parameters is evaluated by the ROC curve.
These findings showed that the three-lncRNA signature might be a novel biomarker for the accurate prognosis prediction of patients with HNSCC.
 lncRNA Blog lncRNA Research and Industry News
lncRNA Blog lncRNA Research and Industry News